Research
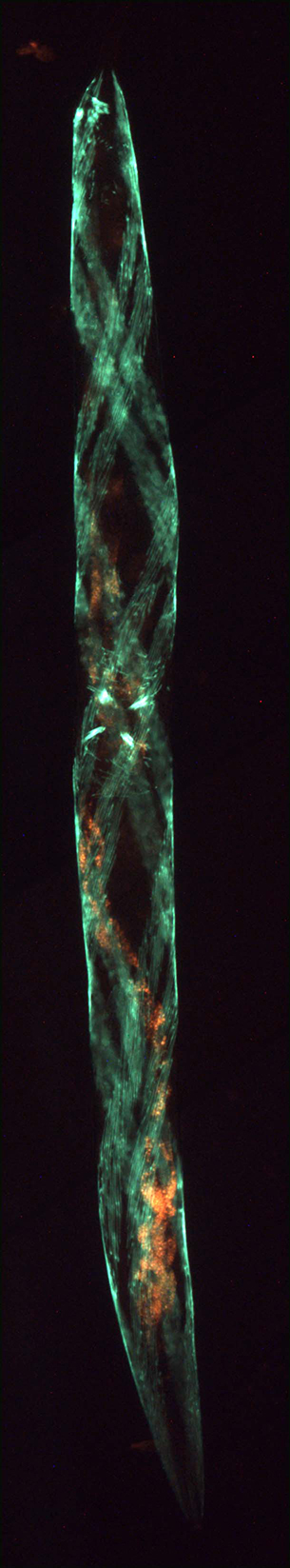
Green fluorescence in the body wall muscles twists around this transgenic worm with a roller phenotype, while the endogenous fluorescence of the intestine appears orange.
Currently the Hope Laboratory is using C. elegans to investigate various human myopathic conditions such as malignant hyperthermia, exertional heat illness, central core disease and late onset axial myopathy ( Baines, K.N., et al. ('17) G3: Genes, Genomes, Genetics, 7, 1451-61.). This is a collaboration with Marie-Anne Shaw of the UK's malignant hyperthermia unit, in the Leeds Institute of Biomedical & Clinical Sciences at St. James's Hospital. We have created strains of C. elegans with the worm's ryanodine receptor modified with single amino acid changes equivalent to those in human ryanodine receptor variants associated with these human conditions. The worm's response to halothane and caffeine changes, with these modifications, in a manner reminiscent of that seen in humans. The ryanodine receptor is an ion channel that allows calcium ions to be released from stores within cells upon excitation of the cell as seen in muscle contraction. Although ryanodine receptors are most abundant within muscle cells, they also function in nerve cells. Our experiments have revealed a possible significance of variant ryanodine receptors for neural diseases.
Previously we have investigated other aspects of muscle cell biology using C. elegans. A collaboration with Elwyn Isaac (also in School of Biology, Leeds) and Netta Cohen (School of Computing, Leeds) involved Systems Biology and Computer Modelling of the C. elegans neuro-muscular system with respect to locomotory behaviour. Mathematical representations of C. elegans movement, across an agar surface and in liquid of varying viscosities, were derived, revealing that C. elegans locomotion from crawling to swimming involves the same gait (Berri, S. et al. ('09) HFSP J., 3, 186-93, Boyle, J. et al. ('11) Frontiers in Behavioral Neuroscience, 5, 10.). In another project, the distinct turnover rates of structural proteins integral to muscle contraction was documented (Ghosh, S. and Hope, I.A. ('10) Eur. J. Cell Biol. 89, 437-48.) A collaboration with Steve Sait (also in School of Biology, Leeds) revealed a genetic link between cold tolerance and longevity in C. elegans (Savory, F. et al. ('11) PLoS One, 6, e24550.; Savory, F.R. et al. ('14) Ecol. Evol., 4, 1176-85.).
Through many years the Hope Laboratory had used a series of approaches, with progressive improvements in efficiency and quality, for determination of the developmental expression patterns for genes across the C. elegans genome. In each approach regulatory regions of C. elegans genes were fused to a reporter gene and transgenic C. elegans lines, transformed with these constructions, were examined to reveal gene expression patterns in situ. The final strategy pursued involved recombineering of fosmids to introduce reporter genes seamlessly into precise positions within genes within large genomic DNA fragments, to maximize the likelihood that all regulatory elements acting on the target gene, endogenously, have been retained (Dolphin, C.T. and Hope, I.A. ('06) Nucleic Acids Research, 34, e72.), in a collaboration with Colin Dolphin (King's College, London). Recombineering was used specifically to examine the operon based expression of the C. elegans sirtuin gene sir-2.1 (Bamps, S. et al. ('09) Mech. Aging Development, 130, 762-70.) and the regulation of transcription factor genes expressed in the nervous system (Bamps, S. et al. ('11) Mol. Genetics Genomics, 286, 95-107, Feng, H. et al. ('12) Gene, 494, 73-84.). Earlier, MultiSite Gateway Recombination had been applied to fuse a reporter gene to promoter regions, from the promoterome, in a collaboration led by Marc Vidal (CCSB, DFCI, Harvard), and including Marian Walhout (UMass Med School). Transgenic C. elegans lines containing these reporter gene fusions were made by microprojectile bombardment rather than the microinjection procedures used previously (Reece-Hoyes, J.S. et al. ('07) BMC Genomics, 8, 27, Dupuy, D., et al. ('07) Nature Biotechnology, 25, 663-8.). Results of these studies contributed to our understanding of how gene expression is orchestrated,genome wide, through development (Craig et al. ('13) BMC Genomics, 14, 249.).
The Hope Laboratory has generated more than four thousand transgenic strains, bearing reporter gene fusions, which have been provided for use in research in other C. elegans laboratories around the world. All the expression pattern data we have generated are available in our database. Our data may also be found in the C. elegans database WormBase, where they are integrated with the genetic, sequence and other data for this organism.
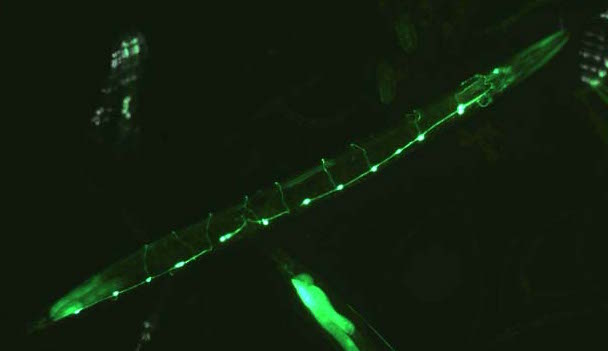
UNC-30::GFP is expressed in type D GABA-ergic motor neurons, with cell bodies and axons in the ventral nerve cord and commissures running round the body (Reece-Hoyes, J.S. et al. ('07) BMC Genomics, 8, 27).
